Kinase and consists of two chains and a bound ligand (search for The protein, in case you're interested, is the SH2 domain of a tyrosine Under Windows you may need to right-click on the You will then get the usual dialogue box asking where Handy hint: To download the above file don't justĬlick on the above link. PDB file (1a07.pdb, Note it is zero not the letter O) These coordinates are the same as those used in the RasMol example. The following link contains the 3D coordinates of a protein structure (in PDB format). 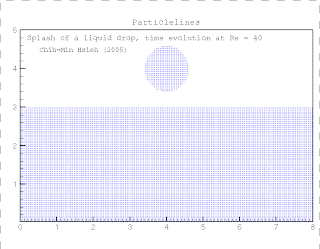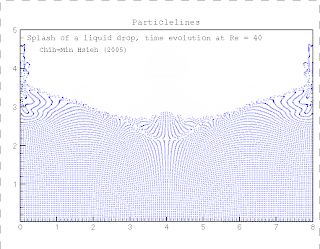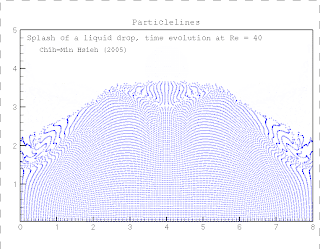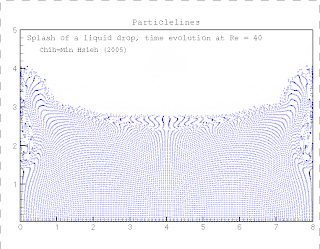
You should note that both JSmol and Rasmol work in very similar ways. To help familarise you with this program, we will go through a few basic commands. You may also want to do a Browser Check (found as a Instructions can also be found under the help Go to the Jmol download page where the link to download the latest version of Jmol can be found.Īn extensive operating manual can be found by clicking on the documentation andįound on the Jmol home page. In the heading, you will find a number of options, including Download. You will need to use it (sometimes another graphics program of at least equalĬapacity will do) to answer some of the courseswork questions and in your project work.Ĭan be obtained from the following link to the The rest of this page gives you instructions for installing Jmol on your computer. Want to make your own images you will have to install Jmol on your own computer. JSmol should work for the images that are embedded within our web pages on most browsers and platforms including on Android and iPbone/iPad. Rasmol, Jmol and JSmol allow users to rotate protein structures, zoom in on them, render them in different ways and using various colouring schemes, labelĪtoms and residues, and so on, as well as exporting images of structures in a number of graphical formats. Which was written in the 90's and was the first program to make fast molecular graphics of complex proteins available on ordinary desktop machines. Java-based program called Jmol which avoids the need for Java when displaying structures embedded in webpages. The principal program that we will be using throughout the PPS course is JSmol which is freely available. Understanding of a protein's structure is to manipulate it interactively using a molecular graphics program. Proteins, by their nature, have complex 3D structures which cannot always be easily appreciated from a 2D picture. Written by Helen White updated by Nick Keep and Clare Sansom JSmol and Jmol JSmol and Jmol: molecular graphics viewers
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
